Toni Balcean turned 101 in September. How’d she beat a century? Simple. “Clean living and good Italian wine.” Case closed! Unless, of course, you like science. A retooled Archon Genomics X PRIZE aims to help scientists better understand healthy aging by sequencing 100 healthy centenarian genomes—in a month, with an accuracy of one error per million base pairs, and for under $1,000 per genome.
All this may sound eerily familiar. In fact, the Archon Genomics X PRIZE was first proposed back in 2006. Singularity Hub covered it in 2008 when the goal was to sequence 100 human genomes in 10 days for less than $10,000 per genome.
Grant Campany, Senior Director of the Archon Genomics X PRIZE, recently told Singularity Hub, “From 2006 to 2009, competitors registered for the Archon Genomics X PRIZE with the best of intentions, but over the past few years the industry has fragmented significantly, so we needed to restructure the competition to be more inclusive of the emerging and established sequencing platforms.”
So, in October 2011, the X PRIZE announced a new set of criteria. The payout remains $10 million; however, the Foundation upped the sequencing period to 30 days and made the target cost $1,000 to reflect rapidly declining sequencing prices.
Most importantly, the reworked competition requires teams produce “medical grade genomes”—a level of accuracy and completeness organizers say doesn’t currently exist.
Our most thorough sequencers leave an estimated 5% to 8% of the typical genome unread. The X PRIZE requires 98% completeness at no more than one error per million base pairs (down from one error per one hundred thousand base pairs previously).
The guidelines also require teams provide haplotype phasing—the mapping of genetic variation between chromosome pairs. Haplotypes are important because they could be closely linked to disease.
But how do you fairly judge the submitted genomes? How do you compare them to a reference genome? This is a major challenge—not only for the competition, but for the field at large.
To solve the problem, Campany says, “We are creating the world’s first open source validation software capable of scoring whole human genome sequences against our quality criteria, namely accuracy, completeness, and haplotype phasing.”
The X PRIZE partnered with EdgeBio to develop this software, dubbed the “validation protocol,” and released it to the community earlier this year for a thorough six month round of testing and improvement. They expect the software to be ready and running come the competition next fall.
There is the purely technological side (the how of it all) and then there is the scientific side (what to study). After mulling a number of possible groups, organizers hit on centenarians—a group who’ve lived through a century of major techological change (see here for their favorite inventions) and add a human element to the competition.
Campany explains, “The older an individual is, the more likely their longevity is linked to their genes. So, we’re recruiting a number of vital supercentenarians [individuals over 110] to provide a source of DNA for teams to sequence as part of the competition. The resulting genomic data will be deposited into dbGaP [the database of Genotypes and Phenotypes] and will be the most highly sequenced and ‘validated’ genomic data ever produced. One of our volunteers, Besse Cooper (116), is the oldest living person in the world, so we are hoping we will be able to answer one key question: What enables these unique individuals to live so long and evade many debilitating diseases?”
The revamped Archon Genomics X PRIZE is exciting, to be sure, but it’ll be a tough competition to win. So, who’s up to the task?
Organizers aim to attract up to five competitors by the May 2013 deadline. So far, two teams are officially in. The first to sign up was Ion Torrent (by Life Technologies) in July. The firm’s Ion Proton Sequencer—sequel to its popular Ion PGM Sequencer—uses semiconductor technology instead of light to sequence genomes. “It’s Watson meets Moore,” the team says.
Learn more about the Ion Proton Sequencer here:
Harvard’s Wyss Institute, under the leadership of Dr. George Church, became the latest entrant in October. Here’s a peek at what the Wyss Institute is all about:
The competition will continue recruiting centenarians through December 2012. They’ve got a pool of 89 on their website and an additional 75 or so through an international centenarian consortium set up by the X PRIZE last fall. In 2013, they will select the final 100 genomes and announce their selections in March. The head-to-head competition is set to run from September 5 to October 4, 2013.
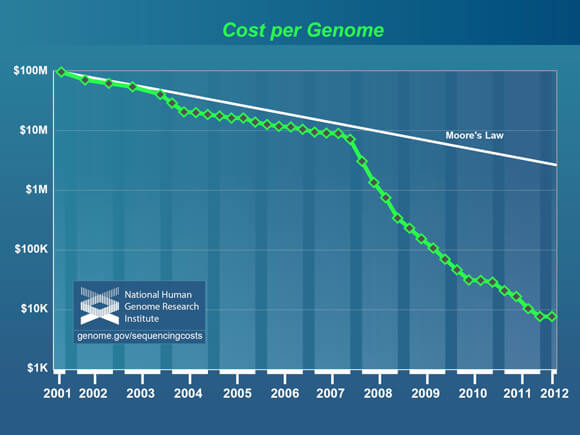
The first fully mapped human genome cost $2.7 billion. A huge sum. But ever since, the cost of genomic mapping has followed a smooth Moore’s Law-like decline, and since 2007 it has vastly outstripped even Moore’s Law.
Crowning an Archon Genomics X PRIZE winner would mean the field has achieved another ten-fold decrease, from $10,000 to $1,000 per sequenced genome—with unmatched accuracy, speed, and completeness. But as much as falling costs are inspiring, it’s that last bit that may be most significant.
In Campany’s words, “We’re trying to catalyze the transformation of genomics from research to clinical use. Challenging teams to achieve our criteria for a medical grade genome is really the centerpiece of the competition—not only to define what that is, but to validate it.“