Genome Study Reveals New Links to Parkinson’s

Share

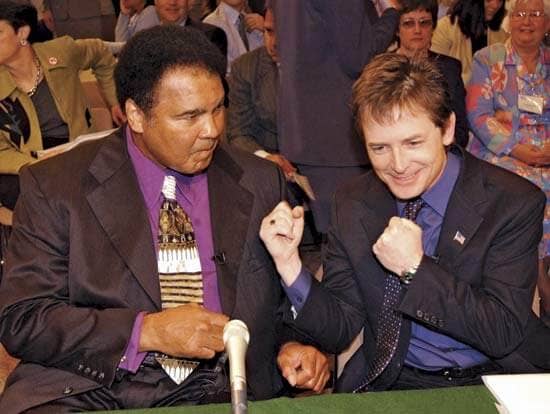
Affordable genetic testing continues to enable scientists to find exciting new discoveries that may help doctors predict, prevent, and treat disease. Two teams of researchers recently published in Nature Genetics (a Japanese team from Kobe University, and a US team from NIH) have collaborated to find that five important genetic variants are linked to Parkinson's. This debilitating brain disease degrades muscle control through a reduction in brain chemicals and affects 1-2% of those over 65. This research was the largest case of genetic testing for Parkinson's, ever. With the amount of genetic data that can now be processed quickly and cheaply, studies like these are just the beginning.
These two Genome Wide Association Studies (GWAS) rely on finding important comparisons of single nucleotide polymorphisms (SNPs). 23andMe declared war on Parkinson's by analyzing SNPs one individual at a time and hope to gather 10,000+ samples total. These two GWAS, however, have already examined the genetics of many thousands of volunteers. By sifting through this massive amount of data, scientists can glean which genetic markers may indicate increased risks of the disease. It's only been in the last few years that genetic testing has been cheap enough to facilitate such studies. As whole genome sequencing becomes cheaper, researchers will be able to study more DNA than just SNPs. This may lead to an even better understanding of the links between genes and illness. We live in a very exciting time - there is an ocean of data in our DNA that is going to be explored in the next few years.
The Japanese team, led by Dr. Tasishi Toda of Kobe University, discovered four important loci for Parkinson's named PARK16, BST, SNCA, and LRRK2. Information was taken from 2011 patients with the disease and 18,381 healthy individuals, all of Japanese descent. In the US, Dr. Andrew Singleton of the NIH led researchers to confirm the SNCA locus and add a new one: MAPT. Their sampling consisted of 1713 patients, and 3978 healthy volunteers followed by 3361 patients and 4573 volunteers, all of European ancestory.
Be Part of the Future
Sign up to receive top stories about groundbreaking technologies and visionary thinkers from SingularityHub.


Why am I giving you all these numbers? To prove a point. We are talking about tens of thousands (30k+) of people each having their genome analyzed on the cheap. That's remarkable, and something that would have been unheard of just five years ago.
The two teams confirmed that PARK16, SNCA, and LRRK2 are important loci for all Parkinson's patients. Interestingly, MAPT was only an indicator for European ancestry, and likewise with BST for Japanese ancestry.
As SNP genetic analysis gets even cheaper, and whole genome studies fall within the budget of research institutions, we are going to see more and more studies like these two. Already individuals can gain access to affordable SNP analysis, and soon the same will be true for whole genome sequencing. That means that you will be able to know for yourself which important markers you carry that may indicate a predilection to an illness. Armed with such knowledge we will all be able to take better control of our health. Hang in there everybody, we're making progress. (Not so) slowly but surely.
[photo credit: Kristen Ryder]
Related Articles
Toxic Clumps in Huntington’s Disease May Protect the Brain Too

AI Can Now Design and Run Thousands of Experiments Without Human Hands. We Aren’t Ready for the Risk to Biosecurity.

Three Countries Own the Lithium Market. An MIT Startup Wants to Break Their Grip.
What we’re reading