NIH Awards $40 Million to Map the Wiring of the Brain

Share
You may never have heard of the Connectome, but chances are it defines nearly everything about who you are. Just as our genome is the totality of our genes, the connectome is the totality of the connections in our brain and nervous system. Many scientists believe that our memories, our personalities, our thoughts are encoded in those connections. We have 100+ billion neurons that link together trillions and trillions of times. The connectome is a chaotic tangle of synapses that seems nearly endless in complexity. The US National Institute of Health wants to map it. Back in 2009 they set up the Human Connectome Project a $60 million dollar endeavor to discover how the human brain is wired. They have recently announced how nearly $40 million of that money will be spent. The short answer: magnets. Researchers at the University of Minnesota, Washington University, Mass General Hospital, Harvard, and UCLA will be mapping the human mind with various instruments, foremost among them the MRI. Along with Siemens Medical Systems they will even create a new magnetic resonance imaging platform, the Connectome Scanner, to peer into the brain with detail no one has seen before. The Connectome is a tangled weave, but it looks like we're on the road to unravel it.
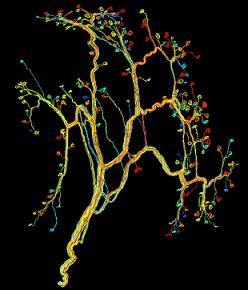
To get a handle on how complex a connectome really is, we should consider how few have actually been mapped. The first and only connectome that is known completely was for a simple worm, C. elegans, that contained just 302 nerve cells. Human brains have hundreds of billions of neurons. Most researchers currently work with small portions of retina and muscle nerves. The complex network shown above is for a very small portion of the muscle tissue near the ear of a mouse. Sebastian Seung, a physicist and neuroscientist at MIT, discusses the absurd complexity of the human connectome in his TED presentation seen in the video below.
Seung's Lab and their colleagues at MIT, Harvard, and around the world have focused on slicing the human brain and microscopically analyzing what they see. They cut open the computer and trace the wires, so to speak, and they do it in three dimensions. Manually, this process is tedious, requiring dozens of researchers double checking each other's work. With enough slices, enough samples, and with artificial intelligence programs that could scan quicker and better than the human eye, this approach to mapping the connectome would work. But it would likely take decades.
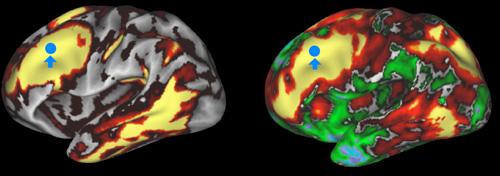
The NIH funded work is taking a different route. MRI scanning may not be able to trace the path of each individual nerve, but it can still tell us a huge amount about how the brain is wired. Teams of researchers at the Washington University and the University of Minnesota Twin Cities will use MRI and fMRI scans to study the brains of 1200 adults - twins and their non twin siblings. These scans will be of greater strength than many typical scans, ranging from 3 Teslas up to 7 or 10 Teslas. Hopefully these scans will grant new insights into how regions of the brain are wired together. The Washington and Minnesota team will also use EEG and MEG to compose movies of brain activity to accompany the MRI scans into the brain's wiring. All of this data will be analyzed by supercomputer and compiled into a Connectome Database Neuroinformatics Platform which will be widely available for neuroscientists to use in the future. Funding for this portion of the Human Connectome Project amounts to around $30 million over five years.
Be Part of the Future
Sign up to receive top stories about groundbreaking technologies and visionary thinkers from SingularityHub.


Another team working at MGH, Harvard, and UCLA will be optimizing MRI scanning. In collaboration with Siemens Medical, this team will be developing a new Connectome Scanner, which will provide greater resolution than traditional techniques (reportedly 4 to 8 times more powerful). The MGH/Harvard/UCLA group's scanner will use diffusion spectrum imaging (DSI), which can pick out the long connections between neurons with greater contrast. The NIH touts DSI as the means to get better data about the connectome and with less time spent in each scan. DSI may be able to give insight into relatively simple bundles of neurons (tens or hundreds of thousands at a time) and how they are linked acorss the length of the brain. This portion of the project will receive $8.5 million in funding.
As massive as the Human Connectome Project is, it is just part of NIH's larger Neuroscience Blueprint, which hopes to develop the tools needed to tackle their three grand challenges: mapping the brain, understanding the neurological basis of pain disorders, and developing therapies for neurological diseases.
Realistically, I don't believe that MRI mapping alone will give us a complete understanding of the human connectome. At some point we're going to need to go in with microscopes and look at slices of the brain. Ultimately such an exploration will need to be automated, as Seung alluded in his talk. The Human Connectome Project and its funding of enhanced MRI, however, will undoubtedly yield valuable insights into our brains. Those insights may indeed lead to better understanding of pain disorders, and finding cures for neurological diseases. It may give us a better idea of how our thoughts and memories are hardwired into the connections in our brains. One day it may even provide the basis for translating the information in our brains to a digital medium. The essence of who we are is likely to be found in the connectome. Once we understand how it is formed, we can start working on improving and changing our minds.
[image credits: Lu et al PLoS Bio 2009, David Van Essen/Washington University, Van J. Wedeen/MGH/Harvard U.]
[video credit: TED]
[source: NIH News, Lu et al PLoS Bio 2009]
Related Articles

An AI Solution to an 80‑Year‑Old Problem Has Shocked Mathematicians

AI Lab Partners Are Rewiring the Hunt for New Drugs

In the Scramble to Power AI, Investors Bet $140 Million on Data Centers at Sea
What we’re reading