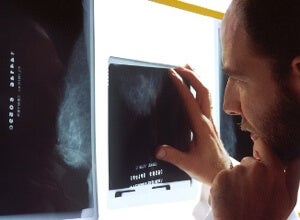
A new study shows that breast cancer can be divided clearly into four different subtypes based on genetic differences. Having been established years ago, the four subtypes are nothing new. What is new is the robust delineation between the groups enabled by an exceptionally large data set. And the unprecedented level of detail uncovered similarities in one group with a treatable form of ovarian cancer, giving hope that the particular breast cancer subtype could one day be eradicated with a similar treatment.
The study is the result of a collaborative effort between 348 researchers. They analyzed the genomes of 825 patients with breast cancers advanced to different stages. But they didn’t just look at genetic sequence, they looked at the genetic switches that turn genes on and off.
One of the breast cancer subtypes, called triple negative breast cancer, was found to be genetically similar to a form of ovarian cancer that can be treated effectively. Computational analysis suggested that, as with the ovarian cancer, the breast cancer might be susceptible to cutting off its blood supply and to some compounds involved in repairing DNA. Any hints toward treating triple negative is sorely needed, as this form of breast cancer is aggressive and has high rates of mortality. Chemotherapy is effective but rates of relapse are high.
The study was published Sept. 23 in Nature.
The researchers are all part of a larger project called the Cancer Genome Atlas that seeks to create an atlas of genetic cancer-causing changes for 20 different types of cancer. The consortium makes their data freely available for use by researchers around the world. If the current study is any indication, the project could lead to important further insights into the subtle genetic differences that exist within a given type of cancer, with the hope that subtle knowledge leads to subtle approaches in finding a cure.
That different genetic subtypes of breast cancer exist is not new knowledge – researchers have known this for decades. However, the current study provides the largest data set yet with which to pick apart those differences. The study also gives ammunition to breast cancer experts who have long thought that cancers ought to be defined based on genetic profiles instead of anatomical location. As Dr. Jay Brooks, chairman of the department of hematology and oncology at Ochsner Clinic Foundation and Hospital in Baton Rouge, LA, pointed out to ABC News, “For a lot of physicians, breast cancer is breast cancer is breast cancer. And the problem is, it isn’t. This is the future of the way we will identify malignancies and the best sort of therapy.”
Clinicians caution, however, that it will be years before any kind of specialized treatments for breast cancer can be developed. Just as the data needed to identify the four subcategories was substantial, so too must the amount of clinical trial data be large in order to show that a new treatment works.
In 2011, breast cancer killed an estimated 40,000 women in the US, according to the American Cancer Society. The same kind of large data analysis performed in the current study could be used to break down other types of cancer into finer genetic subcategories. It’s the subtle kind of approach that is the basis of personalized medicine – tailoring health care according to a person’s unique genetic makeup.
The current acquisition of Complete Genomics by the Chinese company BGI-Shenzhen joins two international genomics powerhouses. Whole genome sequencing – that is, sequencing the ‘other’ 99 percent of the genome that stretches between genes – is Complete Genomics’ specialty. Prior to the acquisition, Complete’s CEO Cliff Reid expressed the company’s intent to be a major player in the future of clinical genomics where patients’ genome sequences will become an integral part of their routine healthcare. Large, high-throughput companies like Complete Genomics and BGI-Shenzhen could provide the massive amount of raw sequence data necessary to identify genetic subgroups within a disease. Tackling cancer, Reid said, is one of Complete’s top goals. As evidenced by the Cancer Genome Atlas, he’s not alone.
The Cancer Genome Atlas represents a breakthrough, not just as far as the current study, but in their approach. Only by pooling the resources of 348 researchers were they able to achieve such a comprehensive and detailed analysis, and then making the data freely available. In doing so they have moved breast cancer research forward. It begs the question, however, why more researchers aren’t banding together in similar ways. Certainly other genetic conditions such as diabetes and cardiovascular disease could benefit from as thorough an analysis. One would think, though, that as genome sequencing, synthesis, and analytical tools become faster, better, cheaper and more widespread, these other types of Atlases will become more the rule than the exception.
[images: Cancer Genome Atlas]
images: Cancer Genome Atlas



