Biotech Startup uBiome Aims To Sequence The Bacteria That Call Our Bodies Home

Share
When you look at your body in the mirror, most of what you consider to be "you" actually isn't you, at least not in a biological sense. That's because there are approximately 10 bacterial cells for every single human cell in the body. Startling, yes, but that's just the tip of the iceberg. Each human body may contain hundreds of thousands of species of bacteria, providing over 350 times the number of genes that is within our own genome, according to an article from Scientific American published last June.
As we consider the issues of health and longevity, the big questions that naturally arise are, what exactly are all these bacteria and what relationship does each have with human physiology?
That's not exactly an easy problem, but fortunately efforts to more rigorously study the bacteria in the body, also known as the microbiome, are underway in research laboratories around the world. Now, a startup called uBiome has recently launched a project on Indiegogo to map human microbiomes through crowdsourced funding and open up sequencing to a much larger pool of people.
With a minimum pledge of $79 to the project, users will receive a kit to obtain samples from their body in order to have their microbiome sequenced. Now, the kit comes with cotton swabs to remove samples of bacterial flora from the mouth, ear, nose, genitalia, or GI tract. Users submit the kit and complete a survey about themselves, and once sequenced, uBiome provides the results, an analysis of what the data mean, charts that compare the user's microbiome to others in the database, and finally the latest research about the bacteria identified.
In the press release, cofounder Zachary Apte said, "We have two aims with uBiome. First, we want to make the science and the technology available to everyone. Now anyone can have their microbiome sequenced. Second, we want to curate the world’s largest microbiome dataset. Citizen science is the answer."
Here's the team's crowdfunding pitch:
With an initial target of 1,000 users, the project so far has nearly 600 backers who have donated enough to receive at least one GI microbiome kit. To receive kits that will enable sequencing at all five different sites, a pledge of $335 is required, and few have opted for the level of mapping.
Though dependent on large numbers of participants to be meaningful, the kind of gross comparative analysis that uBiome provides can be qualitatively insightful. For instance, users could see how their microbioome compares to those in the same geographical region. As the website points out, people who suffer from diabetes or irritable bowel disorder can see how their microflora compare to others with the same conditions. Or perhaps those who are on restricted diets, such as low carb or gluten free, could glean insight into how successful they are with their food choices based on the kinds of bacteria in their GI tracts. Finally, there's also the issue of excesses, like alcohol or coffee, that could show users how their choices are impacting the microecosystems within their bodies.
Be Part of the Future
Sign up to receive top stories about groundbreaking technologies and visionary thinkers from SingularityHub.


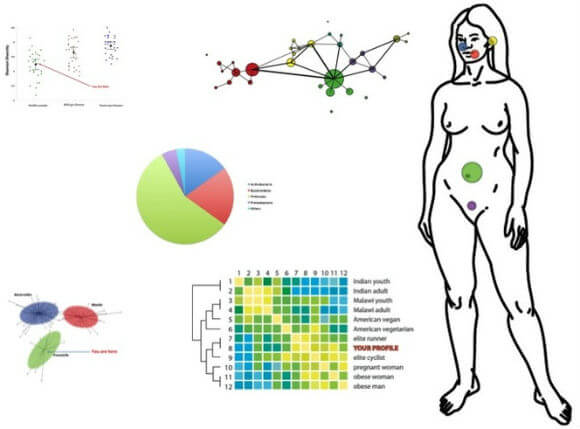
Analysis of microbiome data could give users insight about what's going on inside their bodies that cannot be measured by current medical practices.
Whether any of this information will be able to be tied to specific research that suggests a lifestyle change is in order remains to be seen. But as the human microbiome continues to be researched, more discoveries are bound to make this personal data meaningful.
“We believe the biological information era is going to follow the same trend that the internet did," said uBiome cofounder Jessica Richman in the press release. "When citizens became empowered to explore the internet via search engines like Google, usage skyrocketed. With uBiome, people can explore their personal metagenome from home."
It's a bit overwhelming to realize just how little is understood still about the human body. In fact, it was only a few years ago that doctors at Duke University Medical School proposed a beneficial function of the appendix, long thought of as a vestigial organ left over from evolutionary development. Their theory is that the appendix replenishes good bacteria into the gut, which is vital after things like diarrhea effectively slough the top layer of cells from the GI tract, including bacteria that help to digest consumed food. Without an appendix, a person's GI flora repopulate more slowly, and it's unclear what effects that has on health.
Crowdsourcing the mapping of human microflora can be exactly the kind of complement that researchers need to advance the understanding of this complex problem. Sequencing all of the bacteria in the human body is a monumental task, akin to the Human Genome Project and its challenges. Since 2007, the NIH has been overseeing the Human Microbiome Project, an effort to sequence the microbiomes of 250 volunteers by 200 scientists worldwide. To do this, new kinds of data analysis methods are being developed, and it would be useful to test these against a larger, independent pool that uBiome could acquire.
Understanding how bacterial flora relates to human health will likely be a diverse, multidecade endeavor that is full of surprises. Toward that effort, uBiome's approach offers another way for people to both learn more about their bodies and contribute to important studies.
Perhaps one day, we'll be as equally (if not more) concerned about what the 90 percent of the non-human cells in our bodies are up to as we are about our own genomes. It seems fair since we are providing them room and board, after all.
[image: CIAT International Center for Tropical Agriculture/flickr]
David started writing for Singularity Hub in 2011 and served as editor-in-chief of the site from 2014 to 2017 and SU vice president of faculty, content, and curriculum from 2017 to 2019. His interests cover digital education, publishing, and media, but he'll always be a chemist at heart.
Related Articles

How Fast Are You Aging? New Genetic Clock May Have the Answer

Photosynthetic Drops Soothe Dry Eyes With Sunlight
A Revolutionary Cancer Treatment Could Transform Autoimmune Disease
What we’re reading
