Autism Diagnosed with a Fifteen Minute Brain Scan (Video)
Share
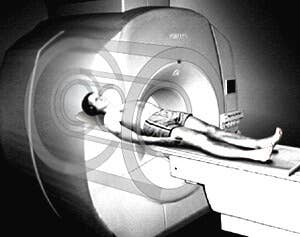
A new technique developed at King’s College London uses a fifteen minute MRI scan to diagnose autism spectrum disorder (ASD). The scan is used to analyze the structure of grey matter in the brain, and tests have shown that it can identify individuals already diagnosed with autism with 90% accuracy. The research could change the way that autism is diagnosed – including screening children for the disorder at a young age.
Currently, ASD is diagnosed behaviorally. There is a list of potential symptoms for autism in the DSM, and anyone exhibiting a certain number of these symptoms can be diagnosed. It’s up to parents or family members to recognize signs of ASD, and to have the child observed by a doctor – only then are they diagnosed. If MRI scans can be shown to rapidly and reliably identify the disorder, it will radically change our diagnostic criteria from behavioral to anatomical.
Researchers at the Institute of Psychiatry (IoP) used MRI scans to evaluate the structure, thickness, and shape of the cerebral cortex (i.e. the outer layer) in subjects’ brains. The study looked at three different groups of males: 20 subjects were healthy controls, 20 had prior diagnoses of ASD, and 19 were diagnosed with ADHD. Each group underwent traditional diagnostic methods first: they were given an IQ test, a psychiatric interview, a physical exam and a blood test. Then the subjects were scanned to see if there were biological correlates to their diagnoses.
The researchers found that ASD patients had special cortical features that allowed their brains to be distinguished from the other two groups. By contrast, ADHD brains could not be distinguished from their healthy counterparts. Because of the small sample size, researchers were unable to distinguish between distinct diagnoses along the autism spectrum (e.g autism vs. Asperger’s). Future research will be needed to determine whether the new technique can tease out diagnostic subcategories of the disorder – and whether the same technique can find these differences in childrens’ brains.
Check out the BBC report:
The disorders of the autism spectrum are poorly understood, to say the least. ASD is largely determined genetically, but the actual mechanisms of its etiology are unclear; it affects the organization of the brain, but in ways we don’t really understand. A 2006 study at Drexel University estimated that a staggering 1 in 170 children have some form of ASD; that figure is more than ten times the estimate given in the 1980’s. Are we diagnosing the disorder more consistently, or is it really on the rise? Again, we don’t know.
Be Part of the Future
Sign up to receive top stories about groundbreaking technologies and visionary thinkers from SingularityHub.


The research is certainly groundbreaking – for the first time, a biomarker has been associated with autism, which has always been diagnosed behaviorally. Still, the work is in its preliminary stages, and won’t be replacing behavioral diagnosis anytime soon (though it will undoubtedly disrupt the DSM-5 revisions of autism’s definition). The test doesn’t give binary “yes or no” results; instead, it plots an individual’s cortical structure in its relative distance from an average non-autistic brain. As Neuroskeptic notes in an excellent summary of the study’s methodology, it might not help diagnosis in borderline cases. As with behavioral symptoms, the anatomical results come in shades of grey – and that still means drawing a diagnostic line somewhere.
The 90% statistic is also misleading, as it indicates the sensitivity of the test rather than the positive predictive value. If we’re asking, “If I have autism, will the brain scan find it?,” the answer is an encouraging 90% “yes.” But if we change the question to “If the scan says I have autism, do I have the ASD?,” that number plummets to something like 5%. In a sobering critique of the study’s hype, Carl Heneghan writes in the Guardian:
Let's think of 10,000 children. Of these 100 (1%) will have autism, 90 of these 100 would have a positive test, 10 are missed as they have a negative test: there's the 90% reported accuracy by the media.
This would seem to limit the research's application to improving diagnosis of suspected individuals, rather than screening the population at large. Still, the research is exciting, even without the hype: it represents the first time that a neuroanatomical difference has been identified in ASD, and that’s an exciting step towards better understanding a baffling disorder. More work lies ahead: the study needs to be replicated in women, and it remains to be seen whether cortical differences can be distinguished in young (i.e. developing) brains.
The statistical issues raise some ethical questions about replacing the behavioral diagnosis with a brain scan. What if you were diagnosed with autism based entirely on the structure of your cortex, never having exhibited any of its behavioral symptoms? Even the study’s authors recognize the complexities involved, e.g. research supervisor Dr. Declan Murphey: “Clearly the ethical implications of scanning people who may not suspect they have autism needs to be handled carefully and sensitively as this technique becomes part of clinical practice.” If you have the brain features, but not the behavioral ones, do you have autism?
These questions will problematize the upcoming DSM-5 and its proposed revisions to ASD's definition. Upcoming studies will undoubtedly put these results to the test, and it remains to be seen what their diagnostic utility will be. But an anatomical element to ASD is exciting news indeed, and should breed a wave of new research into the implications of an identifiable autistic cortex. Hopefully that will pave the way to new treatments for a disorder affecting so many, and one so poorly understood.
[images: DrJerm.com]
Drew Halley is a graduate student researcher in Anthropology and is part of the Social Science Matrix at UC Berkeley. He is a PhD candidate in biological anthropology at UC Berkeley studying the evolution of primate brain development. His undergraduate research looked at the genetics of neurotransmission, human sexuality, and flotation tank sensory deprivation at Penn State University. He also enjoys brewing beer, photography, public science education, and dungeness crab. Drew was recommended for the Science Envoy program by UC Berkeley anthropologist/neuroscientist Terrence Deacon.
Related Articles

Printed Neurons That Mimic Brain Cells Could Slash AI’s Energy Bill
MIT Mined Bacteria for the Next CRISPR—and Found Hundreds of Potential New Tools

The Mad Scramble to Power AI Is Rewiring the US Grid
What we’re reading
