Omri Amirav-Drory wants to be the Bill Gates of the DNA world. Windows revolutionized personal computers by providing a graphic user interface for MS-DOS. Afterwards people didn’t need to be trained in the arcane logic of computer language to be able to use computers.
In an analogous way, Amirav-Drory wants to create a graphic user interface that would empower people to manipulate the arcane logic of DNA. His new software, Genome Compiler (free and available for download at www.genomecompiler.com), converts the various parts of a DNA sequence into easy-to-understand, and easily manipulable, icons. The software turns the complex task of DNA design into an easy drag-and-drop exercise. I caught up with Amirav-Drory recently at Singularity University where he took me through a demo of the program and told me how it might be used by researchers, biology hackers, and what sorts of risks are involved in bringing genetic design into the DIY space.
Trained as a biochemist at Tel Aviv University, Amirav-Drory knows firsthand that designing and constructing the DNA tools used for experiments is labor-intensive and plagued with too many fatal errors. “I hated cloning,” he admits. “It seemed silly to me.”
As a former ‘gene jockey’ myself, I can relate.
To get a sense of the convenience that Genome Compiler could potentially offer requires a general idea of what goes into building a bit of DNA in the lab. Typically, designing an experimental piece of DNA containing the gene you’re interested in starts with manually going through the actual genetic sequence – thousands of A’s, T’s, G’s, and C’s – and, if you were in my lab, using several different colored highlighters to label the important parts on pages in a three-ring binder. The next step would be to design the little stretches of DNA called ‘primers’ to run a polymerase chain reaction (PCR) to amplify your sections of interest. Then you would add enzymes to your amplified DNA to cut out those sections, and later add enzymes to paste them into the newly constructed DNA. Finally, your new little molecular masterpiece is injected into bacteria so that, as the bacteria multiply, so does your DNA. The entire process takes a few days.
And when you find out it didn’t work, you have the exquisite joy of going back and finding out exactly what step went wrong. Like numerous others who have experienced the many trials and many errors of cutting and pasting DNA, Amirav-Drory thinks of the process as “black magic,” and he wants to take the hocus-pocus out of the process.
Genome Compiler is Windows for gene jockeys. Instead of dealing with the actual DNA sequence the user is presented with icons. Drag the E. coli icon into the workspace and the different, specialized parts of its sequence appear as more icons. Icons representing, say, a gene’s start and stop sequences, the promoter regions that control when the gene is expressed, or the region that gives bacteria resistance to antibiotics, can be mixed and matched as easily as rearranging apps on an iPad. The newly designed DNA sequence can be shared with others and, with one click, sent for synthesis.
The actual sequence is, of course, all there behind the colorful, tidy icons. Clicking the icon reveals not only the sequence but information about who sequenced it, the species it was sequenced from, and all the other bookkeeping information researchers are accustomed to. One feature that they’re not used to is the graphic that enables the sequence to be mutated. Click a three base codon, change a letter, and the program informs you that you’re glycine is now a leucine.
And because a sequence created in one lab can be easily shared with others, Amirav-Drory sees Genome Complier as a kind of crowd sourcing tool by which the research community can solve common problems. For instance, he and his friends are already creating a database for expression tags to be placed on different proteins. For example, researchers routinely mutate amino acids to histidines which can then act as a marker to locate, purify and quantify that protein. A critiquable database where researchers can submit their ‘cheat sheets’ of what works best could be a time-saving alternative to searching the literature.
Watch the following video and you’ll see just how user-friendly gene design has become. Graduate students and bio-hackers rejoice.
We don’t often think about the fact that computer data is stored as bits, a stupendous amount of 1’s and 0’s. But that incomprehensible form of information is made comprehensible and useable by graphical interfaces that transform it into words, tools, games. Genome Compiler seeks to do the same for the A’s, T’s, C’s, and G’s of genomic information with its own set of graphical representations. And it’s not only researchers that stand to benefit by sparing their eyes the hours of squinting at endless lines of DNA sequence. Making genes intuitively easy to redesign means researchers won’t be the only ones dabbling in the effects of genetic mutation. With Genome Compiler, DIY biology hackers can become gene jockeys too.
Which is really exciting, but also kind of scary.
Amirav-Drory feels the same way. He says his company, by brining DNA design into the hacker space, is “democratizing the tools of creation.” But what if some unscrupulous bio-hacker wanted to, say, give Borrelia burgdorferi, the bacterium that causes Lyme disease, penicillinase, the gene that destroys penicillin? We’d have a much more virulent form of Lyme disease on our hands. Without giving too many specifics, Amirav-Drory assures me that, not unlike the red flags that go up when people search “how to make a fertilizer bomb” online, safeguards will be put in place to detect such questionable gene retooling.

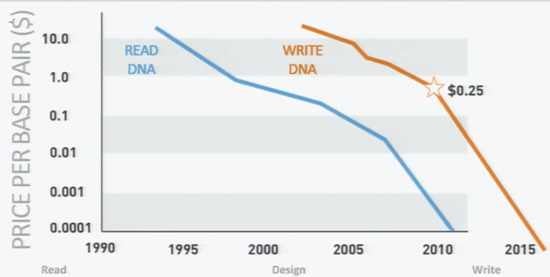
What makes times right – or nearly so – for a program like Gene Compiler is the cost of DNA synthesis.
Amirav-Drory thinks that a program like Genome Compiler is not only useful, but each day becomes increasingly necessary. “There are two technologies that drives this new field of Synthetic Biology: reading DNA and writing DNA. As cost goes down and capability goes up, the need for computer assisted design tools also goes up. Synthetic biology is one of the only technologies that have the potential to scale to the magnitude of our challenges: the need for new ways to produce energy, food, chemicals, plastics, pharmaceuticals and all other commodities our civilization is dependent upon that we currently get from the finite resource of fossil fuels.”
The cost of DNA writing right now is about 25 cents per base pair. At that rate synthesizing an E. coli genome – a relatively small genome of 4.6 million base pairs – would still cost over $1M, too much for the average lab to pay, especially as they typically build up libraries of varied versions of the DNA. But the cost of DNA synthesis is dropping at a super-exponential rate, outpacing even Moore’s Law. Amirav-Drory predicts it will hit 10 cents per base pair this year, and in the near future, when the cost of synthesis drops enough, gone will be the laborious days of manually splicing together pieces of DNA. They won’t be missed.
Amirav-Drory and Genome Compiler are already preparing for such a future. They expect to release a beta version by the end of the year (you can find more information here). Once it’s out there, he thinks that it will ultimately be a tool “democratized” to solve some of society’s biggest problems. “Advanced technology is indistinguishable from nature,” he says. “We’re now learning how to harness this technology. Grey goo? We’re living in the green goo.”
[image credits: Wesolveforx via YouTube, Genome Compiler, Genome.gov, and GREENDIARY]
images: Wesolveforx via YouTube, Genome Compiler, Genome.gov, and GREENDIARY
video: Wesolveforx via YouTube



