Doctors Faced With Rare or Difficult Cancers Can Just ‘Google’ Genetic Treatments
Sequencing cancer genes has become easy and cheap, but information on which drugs might or might not work on particular mutations remains buried in PDF files and in a range of medical journals. So twin researchers Malachi and Obi Griffith, of Washington University in St. Louis, recently launched a drug-gene interaction database that makes the emerging research about as easy to find as a plane reservation on Kayak.com.

Share
Since DNA sequencing began in the 1990s, one of its greatest promises has been that doctors treating cancer could sequence a tumor's DNA, plug the results into a computer and print out a list of the medications most likely to wipe out that particular cancer.
Research into the genetics of cancer is just starting to deliver treatment options two decades later. But twin researchers Malachi and Obi Griffith, of Washington University in St. Louis, recently launched a drug-gene interaction database that makes the emerging research about as easy to find as a flight reservation on Kayak.com.
Genetic information identifying where cancer cells are abnormal can suggest more specific drug treatments than simply labeling the cancer by where in the body it originated.
"Cancers can be broken in very different points in the normal mechanism of how cells live and function. DNA tells us a lot about where the broken parts are, and that’s critical for targeting our fixes. For example, if your car is making a rumbling in the front, you need to know if it’s in the engine or the transmission. And that’s sort of what we don’t know currently," explained U.N.C. Chapel Hill oncologist Neil Hayes in a video about the National Institutes of Health's Cancer Genome Atlas:
Sequencing cancer to find the breaks has become easy and cheap. But information on which drugs might or might not work on particular mutations remains buried in PDF files and scattered across medical journals.
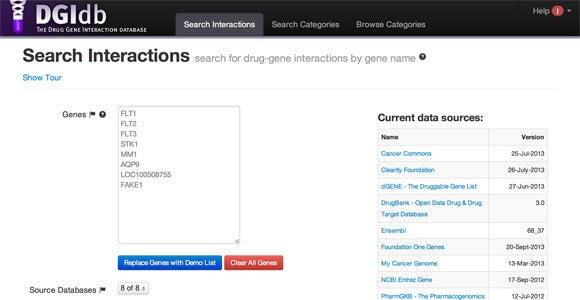
“As we move toward personalized medicine, there’s a lot of interest in knowing whether drugs can target mutated genes in particular patients or in certain diseases, like breast or lung cancer. But there hasn’t been an easy way to find that information,” database co-developer Malachi Griffith said in a news release.
“We wanted to create a comprehensive database that is user-friendly, something along the lines of a Google search engine for disease genes,” he said.
Be Part of the Future
Sign up to receive top stories about groundbreaking technologies and visionary thinkers from SingularityHub.


The Drug Gene Interaction Database isn’t the first searchable database for genes, but it incorporates some advances that bring it much closer to the input/output model imagined as the ultimate benefit of cancer gene sequencing.
It’s specialized for drug interactions, rather than serving as a kind of encyclopedia entry for a particular gene like the database run by the National Center for Biotechnology Information or another run by the Weizmann Institute of Science. It's also largely focused on cancer-related mutations.
The database is geared toward researchers and physician-scientists who want to know whether mutations, likely identified through genome sequencing, could be targeted with existing drug therapies. The database includes drugs approved by the U.S. Food and Drug Administration and others that are in the pipeline but have yet to obtain approval.
DGIdb is also relatively easy to use if you know what you’re looking at: It’s much closer to a Google for genes than to the cumbersome database search queries of yesteryear that still structure many academic and government databases.
Researchers can also run automated queries, such that anytime their research points them to particular gene mutations, they get an automated report on the existing research on those genes. Soon, they will also be able to add their own research to the search engine, pending approval from a moderator.
The Griffiths hope that those features will encourage other researchers to contribute to the database, helping it stay current as genetic research into cancer treatments continues to expand.
“In academia you usually have one grad student that’s tasked with building these. There’s a paper, but [the database] is usually only half usable and it usually dies on the vine because there’s no one to maintain it. We’re trying to change that paradigm,” Obi Griffith told Singularity Hub.
Cameron received degrees in Comparative Literature from Princeton and Cornell universities. He has worked at Mother Jones, SFGate and IDG News Service and been published in California Lawyer and SF Weekly. He lives, predictably, in SF.
Related Articles
Toxic Clumps in Huntington’s Disease May Protect the Brain Too

AI Can Now Design and Run Thousands of Experiments Without Human Hands. We Aren’t Ready for the Risk to Biosecurity.

Three Countries Own the Lithium Market. An MIT Startup Wants to Break Their Grip.
What we’re reading