Scientists first transcribed the genome—or complete genetic code—of a free living organism in 1995. Sequencing the bacterium H. Influenzae took a little over a year, cost about $1 million, and required a (then) significant amount of computing power. In 1997, Escherichia coli (E. coli) similarly surrendered its genetic secrets.
Less than two decades on, the entire genome of E. coli has again been sequenced. This time, however, researchers used a handheld machine the size of a smartphone. Oxford Nanopore’s MinION genetic sequencer plugs into a laptop by USB and taps cloud computers for the heavy lifting.
The study, led by the Ontario Institute for Cancer Research’s Dr. Jared Simpson and recently published in the journal Nature Methods, shows just how much can be done with the tiny device, and the researchers believe more complex reads are possible. How does it work?
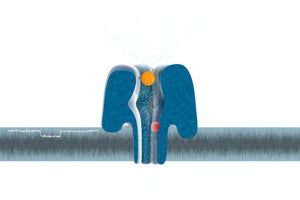
The MinION device’s nanopore technology embeds special proteins in an electrically resistant membrane. DNA molecules are drawn through these protein “gates” and, as they flow through, they disrupt an electric field. Sensors record each molecule’s unique electrical fingerprint and send it on to the central computer for identificantion.
In recent years, the price of human genome sequencing has fallen dramatically—going from hundreds of millions over a decade ago to $1,000 per genome last year. But the machines doing the work are still expensive, large, and confined to laboratories.
MinION is small, of course, but also relatively affordable. Oxford Nanopore expects to offer each device for a retail price of less than $900.
The device’s diminutive size, however, does have its drawbacks—primarily, it is much less accurate than larger, more powerful sequencers. And while showing the potential of the technology was one result of the research, perhaps more significant were the analytical methods and software devised to overcome its weaknesses.
 Using a three-stage process and probabilistic model of each DNA molecule’s electrical signal as it moved through a nanopore, the team was able to overcome the device’s error rate to correct and ultimately refine their final sequence.
Using a three-stage process and probabilistic model of each DNA molecule’s electrical signal as it moved through a nanopore, the team was able to overcome the device’s error rate to correct and ultimately refine their final sequence.
“We were able to mathematically model nanopore sequencing and develop ways to reconstruct complete genomes off this tiny sequencer,” said Simpson.
Even so, assembling the entire E. coli genome, about 4.6 million bases (or pairs of genetic “letters”) took four runs. At more than three billion bases, the human genome is much bigger, and at least for now, far out of reach in terms of cost and practicality.
That said, the group believes the methods they developed do offer the opportunity to scale up to more complex animals, maybe even humans, and in the meantime, there are other just as useful applications for portable genomic sequencers.
One such use, personalized medicine, might make it possible to sequence tumor genes on the spot, allowing doctors to more rapidly develop therapies tailored to the patient. Another use might be in epidemiology—in the event of an outbreak, we could quickly sequence a virus to help develop a defense against it.
“Because the instrument is very portable we have managed to use the instrument in the field, for example sequencing the Ebola virus in Guinea,” said the University of Birmingham’s Nicholas Loman, another of the study’s authors.
The only flaw? Because the machine requires an internet connection and cloud computing capabilities, it can be hard to use in areas with spotty online access. One solution might be to ask the local computer to foot more of the burden. (And more ubiquitous global internet access may be just around the corner too.)
In the foreseeable future, we expect the biggest breakthroughs in decoding and understanding the genome will still come from large studies conducted using significantly bigger machines. But ultimately, when it comes to bringing the power of genomics to the masses? Small will be powerful too.
Image Credit: Oxford Nanopore



