Influenza: The Search for a Universal Vaccine

Share
The current 2017-18 flu season is a bad one. Hospitalization rates are now higher than in recent years at the same point, and infection rates are still rising. The best line of defense is the seasonal influenza vaccine. But H3N2 viruses, like the one that’s infecting many people this year, are particularly hard to defend against, and this year’s shot isn’t very protective against H3N2.
Producing an effective annual flu shot relies on accurately predicting which flu strains are most likely to infect the population in any given season. It requires the coordination of multiple health centers around the globe as the virus travels from region to region. Once epidemiologists settle on target flu strains, vaccine production shifts into high gear; it takes at least six months to generate the more than 140 million doses necessary for the American population.
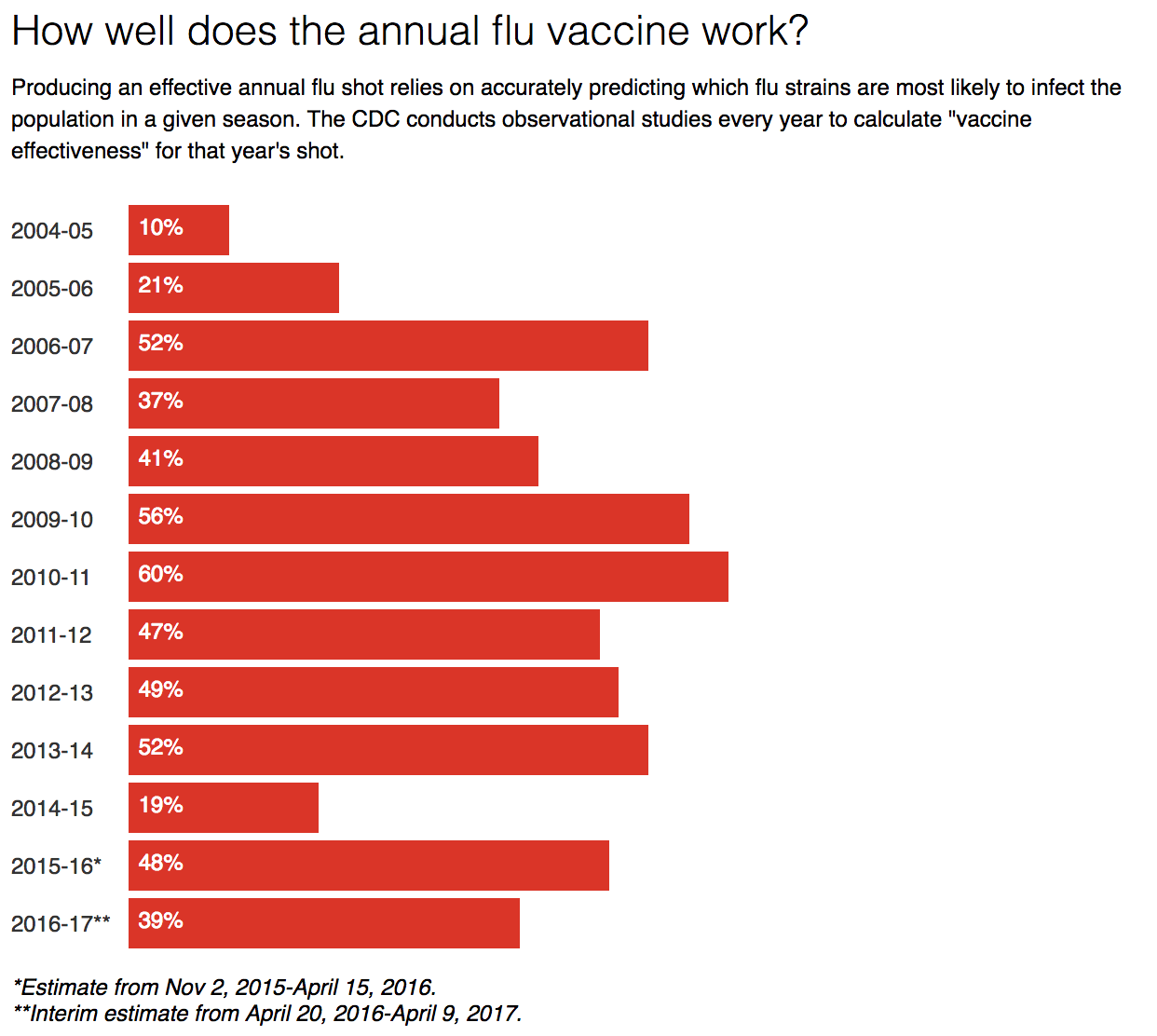
Chart: The Conversation, CC-BY-ND Source: Centers for Disease Control and Prevention
Incorrect or incomplete epidemiological forecasting can have major consequences. In 2009, while manufacturers were preparing vaccines against the forecasted strains, an unanticipated H1N1 influenza virus emerged. The prepared seasonal vaccine didn’t protect against this unanticipated virus, causing worldwide panic and over 18,000 confirmed deaths. This was likely only a fraction of the true number of deaths, estimated to exceed 150,000. Better late than never, a vaccine was eventually produced against the emergent H1N1, requiring a second flu shot that year.
Given that influenza has caused the majority of pandemics over the past 100 years—including the 1918 flu that resulted in as many as 50 million deaths—we’re left with the question: Can scientists produce a “universal” vaccine? An ideal version would be capable of protecting against diverse strains of influenza and wouldn’t require a yearly shot for you.
Vaccines Prime the Immune System to Fight
By the 18th century, and arguably much earlier in history, it was commonly known that a survivor of smallpox would not come down with it again upon subsequent exposure. Somehow, infection conferred immunity against the disease. In fact, people recognized that milkmaids who came into contact with cowpox-ridden cattle would similarly be protected from smallpox.
In the late 1700s, farmer Benjamin Jesty inoculated his family with cowpox, effectively immunizing them against smallpox. Physician Edward Jenner then catapulted humanity into a new age of immunology when he lent scientific credence to the procedure.
So if one inoculation of cowpox or one exposure to (and survival of) smallpox confers a decade’s worth or even lifelong immunity, why are individuals encouraged to receive the flu vaccine every year?
The answer lies in how quickly the influenza virus’s anatomy changes. Each virus consists of a roughly spherical membrane encapsulating constantly mutating genetic material. This membrane is peppered with two types of “spikes”: hemagglutinin, or HA, and neuraminidase, or NA, each made up of a stem and a head. HA and NA help the virus with infection by binding to host cells. They mediate the entry of the virus into the cell and, once it replicates, the eventual exit.
Once a doctor injects a vaccine, an individual’s immune system gets to work by making antibodies that recognize, for example, the hemagglutinin it contains. The next time that hemagglutinin shows up—such as in the form of the virus strains the vaccine mimicked—the body’s immune cells recognize them and fight them off, preventing infection.
For vaccine developers, one frustrating characteristic about influenza’s mutating genome is how rapidly HA and NA change. These constant alterations are what send them back to the drawing board for new vaccines every flu season.
Different Methods to Design a Vaccine
The smallpox vaccine was one of the earliest to use the “empirical paradigm” of vaccinology—the same strategy we largely use today. It relies on a trial-and-error approach to mimic the immunity induced by natural infection.
In other words, vaccine developers believe the body will react to something in the inoculation. But they don’t focus on which specific patch of the virus is causing that immune response. It doesn’t really matter if it’s a reaction to a small patch of HA that many strains share, for instance. When using an entire virus as starting material, it’s possible to get many different antibodies recognizing many different parts of the virus used in the vaccine.
Be Part of the Future
Sign up to receive top stories about groundbreaking technologies and visionary thinkers from SingularityHub.


The seasonal flu shot generally fits into this empirical approach. Each year, epidemiologists forecast which flu strains are most likely to infect populations, typically settling on three or four. Researchers then attenuate or inactivate these strains so they can act as the mimics in that year’s influenza vaccine without giving recipients the flu. The hope is that an individual’s immune system will respond to the vaccine by creating antibodies that target these strains; then when he or she comes into contact with the flu, the antibodies will be waiting to neutralize those strains.
But there’s a different way to design a vaccine. It’s called rational design and represents a potentially game-changing paradigm shift in vaccinology.
The goal is to design some molecule—or immunogen—that can trigger the production of effective antibodies without requiring exposure to the virus. Relative to current vaccines, the engineered immunogen may even allow for more specific responses, meaning the immune response targets particular regions of the virus. There’s the possibility of greater breadth, too, meaning it could target multiple strains or even related viruses.
This strategy works to target specific epitopes, or patches of the virus. Since antibodies work by recognizing structures, the designers want to emphasize to the immune system the structural properties of the immunogens they’ve created. Then researchers can try to design candidate vaccines with those structures in hopes they’ll provoke the immune system to produce relevant antibodies. This path might let them assemble a vaccine that elicits a more effective and efficient immune response than would be possible with the traditional trial-and-error method.
Promising headway has been made in vaccine design for respiratory syncytial virus using this new rational paradigm, but efforts are still underway to use this general approach for influenza.
Toward a Universal Flu Vaccine
In recent years, researchers have isolated a number of potent, infleunza-neutralizing antibodies produced in our bodies. While the antibody response to influenza is primarily directed at the head of the HA spike, several have been found that target HA’s stem. Since the stem is more constant across viral strains than the head, this could be flu’s Achilles’ heel, and antibodies that fix on this region may be a good target for vaccine design.
Researchers are pursuing a number of approaches that could cause the body to produce these antibodies of interest before becoming infected. In one strategy, scientists attached lab-made copies of hemagglutanin stems to a spherical protein nanoparticle. The resultant structure isn’t a virus and doesn’t even contain any viral genetic material. But it looks a lot like a virus to the body’s immune system, and so elicits a good antibody response. And, because only the stem is attached to the nanoparticle, the immune system can focus the antibody response on these regions which are more similar from strain to strain than the head. This general approach has seen success both in mice and ferrets, but further testing is required before it can be tried in people.
With current technology, there may never be a “one and done” flu shot. And epidemiological surveillance will always be necessary. However, it is not inconceivable that we can move from a once-per-year model to a once-every-10-years approach, and the field has been making huge strides to achieve this.![]() This is an updated version of an article originally published on Jan. 11, 2017. This article was originally published on The Conversation. Read the original article.
This is an updated version of an article originally published on Jan. 11, 2017. This article was originally published on The Conversation. Read the original article.
Image Credit: CNK02 / Shutterstock.com
Ian is a PhD student in the Chemical & Physical Biology program. He received his BS in physics from the University of Richmond where he conducted research in a systems biology laboratory studying signal propagation in genetic networks and cellular behavior under stress. His interests, broadly speaking, are in computational structural immunology and systems immunology.
Related Articles

Orbital Airbag Could Shield Earth From Devastating Solar Storms

This Week’s Awesome Tech Stories From Around the Web (Through June 7)
Toxic Clumps in Huntington’s Disease May Protect the Brain Too
What we’re reading