Can We Resurrect Extinct Species? Scientists Put Jurassic Park to the Test

Share
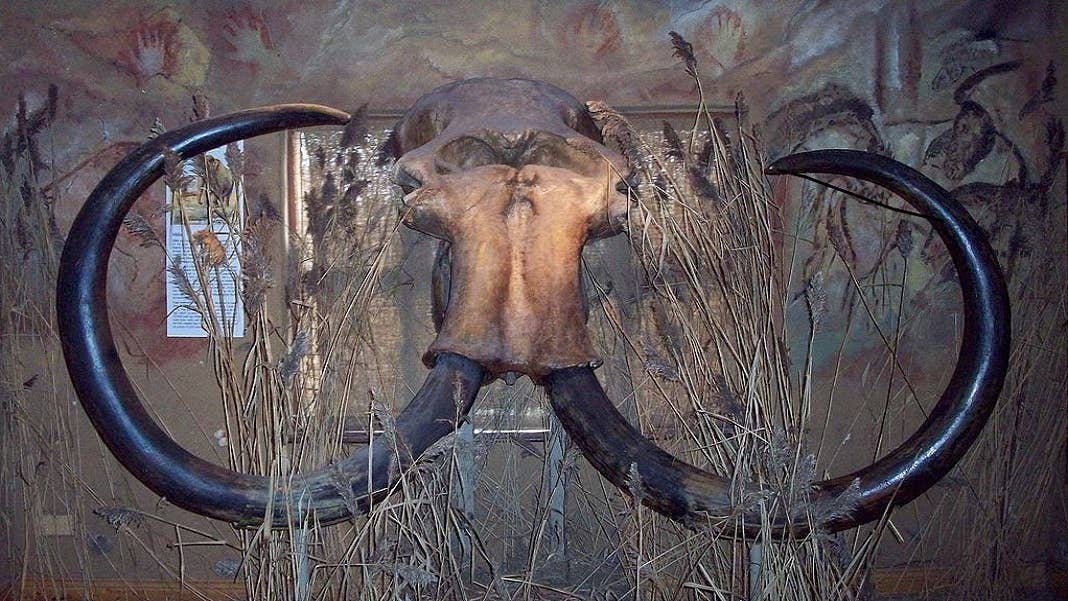
De-extinction grabbed our imagination in the 90s with Jurassic Park. Scientists have since asked: how possible is it?
According to a new study, nearly impossible. But wait—it’s not all bad news. While bringing back a faithful copy of an extinct species may be impossible, we could bring back a hybrid species that’s a genetic mix between an extinct species and its modern descendant.
Published in Current Biology, the study eschews the grandiose mammoth, instead focusing on a tiny test case: the Christmas Island rat. Hefty in size and loudly vocal when invading docked ships and their cargo, the rodents were last seen in the 1900s. With a stroke of luck, the team recovered DNA from two well-preserved museum samples and compared them against a close relative: the Norway brown rat, a popular lab model for genetic studies today.
The two species share roughly 95 percent of their total genome. But the remaining five percent, mostly involved in the immune system and sense of smell—something rats heavily rely on—were “unrecoverable.”
In other words, even if the rats can be brought back, they’ll be significantly changed from the original rat. The results may guide the effort to bring back a “modernized” version of the woolly mammoth from elephants, which have a similar evolutionary distance as Christmas Island rats and Norway brown rats.
"It is very, very clear that we are never going to be able to get all the information to create a perfect recovered form of an extinct species," said Dr. Tom Gilbert, an evolutionary geneticist at the University of Copenhagen who led the study. "There will always be some kind of hybrid."
Genetic De-Extinction
Let’s backtrack. How does de-extinction work?
It comes down to manipulating DNA. One idea is cloning. This requires highly preserved DNA synthesized from scratch. But ancient DNA is usually heavily fragmented, like a historical book that’s been through the shredder. This makes piecing together the old genome—and breeding living animals from it—nearly impossible (sorry kids, the Jurassic Park approach won’t work.)
Another option is to re-write the genome of a modern animal to better match it to its extinct cousin. With the rise of the gene editor CRISPR, this approach “is most likely to apply to the largest number of extinct species,” wrote the team.
The recipe for an ancestral glow-up is relatively simple on paper. The first step is identifying a closely related species. Its genome is then faithfully sequenced at a high resolution. The resulting data is used to construct a reference genome.
Then comes the hard part: finding a DNA sample of the extinct animal. Here, Gilbert’s team got lucky, finding two samples from the skin of Christmas Island rats collected over a century ago. Carefully stored at the Oxford University Museum of Natural History collections, the samples yielded shredded but valuable chunks of DNA.
The team next compared those DNA fragments with the reference genome. The Norway brown rat isn’t an exact descendant of the Christmas Island rat—the two diverged roughly 2.6 million years ago. But on the evolutionary scale, they’re close cousins. Like matching an ancient, broken-down copy of a book to a similar, modern one, the team was able to reconstruct nearly 95 percent of the Christmas Island rat’s genome.
The percentage may seem high, but it’s not perfect. The team scratched their heads and wondered why the last five percent remained a “black box.”
“Every bit of DNA that we could recover, we got,” Gilbert said to New Scientist. “There’s a five percent fraction we can’t make sense of.”
They first ruled out potential technological stumbles and sequencing limits—no luck. They then compared the Christmas Island rat’s genome to that of other modern rats, and an answer emerged. It’s evolution. Some genetic information was lost between the extinct species and their modern counterparts, making it nearly impossible to time-travel back on the genomic scale.
The “black box” parts of the genome weren’t random. Mapping nearly 130,000,000 DNA letters missing from the modern reference, the team realized that nearly a quarter covered key genes. Among those were some that help develop a soft coat and strong nails. Others relate to the sense of smell and pheromones, essential for both a rat’s survival and social behaviors.
Be Part of the Future
Sign up to receive top stories about groundbreaking technologies and visionary thinkers from SingularityHub.


A De-Extinction Conundrum
So what to make of all this?
To Gilbert, the answer is clear: even if it’s theoretically possible to reconstruct the Christmas Island rat by CRISPRing a Norway rat, the result will be Frankenstein-esque. The lab-created hybrids could face tremendous challenges when reintroduced to a modern environment.
“Given the role of olfaction in many critical behaviors,” they wrote, “revived Christmas Island rats could struggle to forage for food, detect predators, or find mates—all behaviors tantamount to survival.”
To the team, however, the point of the study wasn’t to bring back a rat. “We aren’t actually planning to do it, as probably the world doesn’t need any more rats,” joked Gilbert.
Rather, it’s to probe the limits of de-extinction. As a leading expert in the field, Gilbert has been hesitant on its possibility. “All ancient DNA is crap,” he said back in 2017, not completely referring to the genetic material’s quality, but also that some key genes evolve very fast. “Missing genes” eliminated through evolution will always be a problem.
What troubles Gilbert is that the omitted chunks of ancient DNA aren’t arbitrary. Rather, not including them in the reconstruction may severely change an animal’s biology and behavior. If a de-extinct animal behaves differently, especially when introduced into today’s environment—which is hundreds if not millions of years apart from its past habitat—have we just made a facsimile? In other words, are we willing to accept a reconstructed mammoth-like being that genetically amounts to a furry elephant?
His team also acknowledges that comparing a more closely-related species could help better reconstruct the extinct genome. One option is the black rat, which roams our neighborhoods. As a next tentative step—and a proof of concept—the team is considering using CRISPR to edit the black rat’s genome to resemble that of a Christmas Island rat.
To Ben Novak, lead scientist at the non-profit Revive & Restore, which focuses on genetic methods to enhance biodiversity, “anyone pursuing de-extinction has to settle on the fact that we want to get as close as we can to something that fools the environment,” he said to Science News. He plans to apply the study’s analysis to his own work. As the program manager for Biotechnology for Bird Conservation, Novak has long focused on using genetic and cloning techniques for the “Great Passenger Pigeon Comeback.” It’ll be a tough project: the genetic divergence between the passenger pigeon and its modern version is over two times more than that between the Christmas Island and Norway rats.
As for Gilbert, he questions whether our focus should be more on the present than on the past. As a technology, he said, de-extinction is “fascinating.” But with so many animals threatened, “one has to wonder if that's the best use of money as opposed to keeping the things alive that are still here.”
Image Credit: Ogmios/Wikimedia Commons
Looking for ways to stay ahead of the pace of change? Rethink what’s possible. Join a highly curated, exclusive cohort of 80 executives for Singularity’s flagship Executive Program (EP), a five-day, fully immersive leadership transformation program that disrupts existing ways of thinking. Discover a new mindset, toolset and network of fellow futurists committed to finding solutions to the fast pace of change in the world. Click here to learn more and apply today!
Dr. Shelly Xuelai Fan is a neuroscientist-turned-science-writer. She's fascinated with research about the brain, AI, longevity, biotech, and especially their intersection. As a digital nomad, she enjoys exploring new cultures, local foods, and the great outdoors.
Related Articles

How Fast Are You Aging? New Genetic Clock May Have the Answer

Photosynthetic Drops Soothe Dry Eyes With Sunlight
A Revolutionary Cancer Treatment Could Transform Autoimmune Disease
What we’re reading